Manually weighted taxonomy classifiers improve species-specific rumen microbiome analysis compared to unweighted or average weighted taxonomy classifiers
Bioinformatics Bioinformatics MethodologyPosted by mhb on 2025-11-06 09:53:29 |
Share: Facebook | Twitter | Whatsapp | Linkedin Visits: 256
Introduction
This study addresses the challenges in accurately classifying species in the rumen microbiome, a crucial area for ruminant animal nutrition and productivity. Previous methods have struggled with rumen-specific taxonomic resolution due to a lack of suitable reference data. This paper introduces a manually weighted taxonomy classifier (MWTC) designed specifically for the rumen microbiome, combining shotgun and amplicon sequencing data to improve accuracy in species-level classification.
Methods
-
Taxonomy Classifiers: The study compared three classifiers: unweighted taxonomy classifier (UWTC), average weighted taxonomy classifier (AWTC), and manually weighted taxonomy classifier (MWTC).
-
Databases: The classifiers were evaluated using several prominent databases, including SILVA, Greengenes, GTDB, NCBI RefSeq, and RDP, focusing on species-level classification.
-
Datasets: Data from both in vivo and in vitro rumen samples of Hanwoo cattle were used to evaluate classifier performance.
Results
-
Performance Evaluation: MWTC showed significant improvement in classification accuracy, with higher classification counts, fully classified ratios, and reduced error rates compared to UWTC and AWTC, especially at the species level.
-
Database Comparison: NCBI RefSeq and RDP performed best with MWTC, showing up to 47% error reduction at the species level. Other databases like GG2 and SILVA showed less improvement due to inherent limitations in their classifications.
-
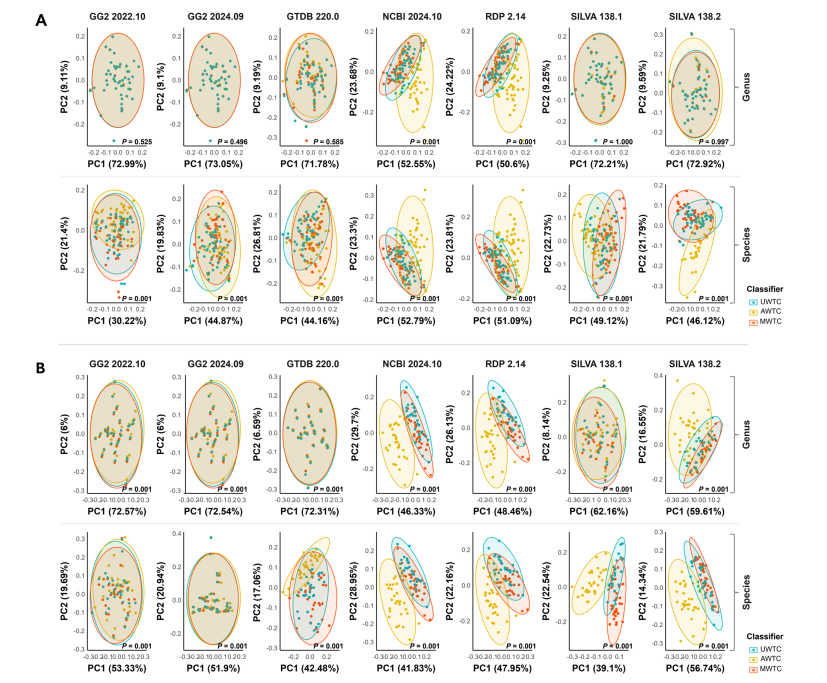
Alpha and Beta Diversity: The MWTC improved diversity metrics, particularly at the species level, across both full-length and V3-V4 amplicon sequences.
Discussion
-
The study emphasizes the importance of breed-specific classifiers for improving microbiome analysis. While databases like GG2 and SILVA struggled with accuracy, NCBI RefSeq and RDP, when paired with MWTC, offered the most reliable results for species-level classification.
-
The findings suggest that integrating rumen-specific taxonomic weights, rather than relying solely on general databases, can substantially enhance microbiome studies.
Conclusion
The study concludes that MWTC, when applied to rumen microbiome analyses, significantly improves species-level classification accuracy. It recommends using NCBI RefSeq for species classification, with MWTC as the preferred method for processing rumen microbiome data. The results underscore the need for breed-specific classifiers in microbiome research.
Search
Categories
Recent News
- Clinical application research on the titanium metal metatarsal prosthesis designed through FEA and manufactured by 3D printing
- Manually weighted taxonomy classifiers improve species-specific rumen microbiome analysis compared to unweighted or average weighted taxonomy classifiers
- A deep learning based smartphone application for early detection of nasopharyngeal carcinoma using endoscopic images
- sCIN: a contrastive learning framework for single-cell multi-omics data integration
- CustOmics: A versatile deep-learning based strategy for multi-omics integration
- Tracking temporal progression of benign bone tumors through X-ray based detection and segmentation
- mCNN-GenEfflux: enhanced predicting Efflux protein and their super families by using generative proteins combined with multiple windows convolution neural networks
- Evaluation of normalized T1 signal intensity obtained using an automated segmentation model in lower leg MRI as a potential imaging biomarker in Charcot– Marie–Tooth disease type 1A
Popular Paper
- sCIN: a contrastive learning framework for single-cell multi-omics data integration
- CustOmics: A versatile deep-learning based strategy for multi-omics integration
- A deep learning based smartphone application for early detection of nasopharyngeal carcinoma using endoscopic images
- Tracking temporal progression of benign bone tumors through X-ray based detection and segmentation
- Clinical application research on the titanium metal metatarsal prosthesis designed through FEA and manufactured by 3D printing