mCNN-GenEfflux: enhanced predicting Efflux protein and their super families by using generative proteins combined with multiple windows convolution neural networks
Bioinformatics Bioinformatics MethodologyPosted by mhb on 2025-11-19 10:55:30 | Last Updated by mhb on 2026-04-18 00:06:55
Share: Facebook | Twitter | Whatsapp | Linkedin Visits: 194
Introduction
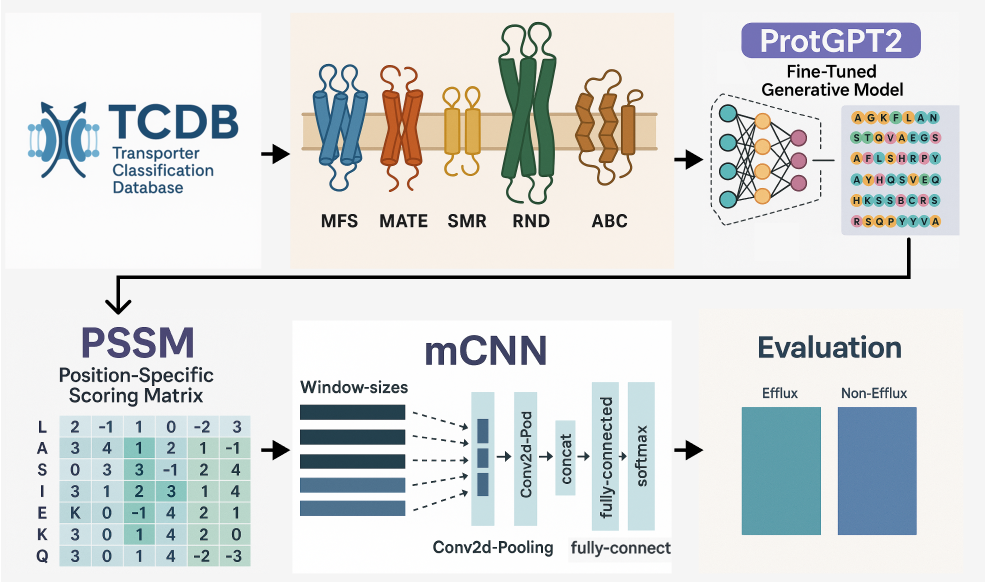
Efflux transporters are essential components of bacterial defense systems because they remove harmful substances, including antibiotics, from the cell. These transporters are divided into five major families: ABC, MFS, MATE, RND, and SMR. Their extensive sequence diversity and limited functional annotation make them difficult to classify using traditional computational approaches. Advances in artificial intelligence, particularly in generative modeling, have opened new possibilities for analyzing and designing biological sequences. Models such as ProtGPT2 can generate realistic protein sequences that mirror natural diversity, allowing researchers to address data scarcity and improve classification performance.
The GenEfflux framework introduces a new approach that combines ProtGPT2-generated efflux protein sequences with a multi-window convolutional neural network. This design captures important structural motifs and evolutionary signals often overlooked by conventional single-feature or alignment-based methods. As a result, the framework offers improved classification accuracy and enhances understanding of efflux protein behavior, which is critical for studying antibiotic resistance and protein regulation.
Method
The GenEfflux approach is built upon two major components: generative sequence expansion and deep feature extraction.
1. Sequence Generation with ProtGPT2
ProtGPT2 is used to generate new efflux protein sequences that closely resemble natural proteins in their amino acid composition. These sequences supplement the original dataset, increasing diversity across the five efflux families. This expanded dataset improves model generalization and reduces problems caused by limited annotated sequences.
2. PSSM-Based Evolutionary Feature Extraction
Position-Specific Scoring Matrices (PSSMs) are computed for both natural and generated sequences. PSSMs capture evolutionary conservation at each sequence position, highlighting functionally important regions. These matrices serve as the input for the classification model and provide richer information than raw sequences alone.
3. Multi-Window Convolutional Neural Network (MCNN)
A multi-window CNN architecture is employed to extract both local and global evolutionary features. By using multiple filter sizes, the network identifies sequence motifs, interaction sites, and patterns associated with efflux activity. This architecture captures relationships that single-window CNNs or alignment-based approaches may miss.
4. Performance Evaluation
The model is evaluated using five-fold cross-validation across three efflux protein classes. Metrics such as sensitivity, specificity, accuracy, MCC, AUC, and F1-score are used to compare GenEfflux with the baseline DeepEfflux model. Statistical significance is assessed using paired t-tests.
Results
The GenEfflux model shows consistent improvements compared with the DeepEfflux baseline across multiple efflux protein classes.
Class B Performance
GenEfflux demonstrates major improvements:
-
Sensitivity increased from 0.5385 to 0.9999.
-
MCC increased from 0.4397 to 0.9327.
These results show a substantial enhancement in detecting positive efflux proteins.
Class C Performance
Significant gains are observed:
-
Accuracy improved from 0.8977 to 0.9668.
-
MCC improved from 0.7668 to 0.9331.
This indicates that GenEfflux provides more reliable classification across both positive and negative classes.
Amino Acid Composition Analysis
Generated sequences were compared to natural sequences across the five efflux families. Measures such as Jensen–Shannon Divergence, cosine similarity, and chi-square tests (all p = 1.0) show strong similarity between synthetic and natural sequences, confirming that ProtGPT2 generates biologically meaningful sequence variations.
Statistical Evaluation
Cross-validation and paired t-tests demonstrate that performance improvements are significant. In Class C, the increase in AUC (0.9805 vs. 0.9789) is statistically significant with p = 0.0076.
Conclusion
The GenEfflux framework presents an effective method for generating and classifying efflux proteins using deep learning techniques. By integrating ProtGPT2-generated sequences with a multi-window convolutional neural network, the approach captures important evolutionary and functional patterns. The results show substantial improvements in accuracy, sensitivity, and MCC across key efflux protein classes. The close similarity between generated and natural sequences further confirms the suitability of ProtGPT2 for expanding protein datasets. Overall, GenEfflux offers a powerful tool for studying efflux transporters, understanding bacterial resistance mechanisms, and supporting future research in protein engineering and computational biology.
Read Full Paper Here
Search
Categories
Recent News
- Clinical application research on the titanium metal metatarsal prosthesis designed through FEA and manufactured by 3D printing
- Manually weighted taxonomy classifiers improve species-specific rumen microbiome analysis compared to unweighted or average weighted taxonomy classifiers
- A deep learning based smartphone application for early detection of nasopharyngeal carcinoma using endoscopic images
- sCIN: a contrastive learning framework for single-cell multi-omics data integration
- CustOmics: A versatile deep-learning based strategy for multi-omics integration
- Tracking temporal progression of benign bone tumors through X-ray based detection and segmentation
- mCNN-GenEfflux: enhanced predicting Efflux protein and their super families by using generative proteins combined with multiple windows convolution neural networks
- Evaluation of normalized T1 signal intensity obtained using an automated segmentation model in lower leg MRI as a potential imaging biomarker in Charcot– Marie–Tooth disease type 1A
Popular Paper
- sCIN: a contrastive learning framework for single-cell multi-omics data integration
- CustOmics: A versatile deep-learning based strategy for multi-omics integration
- A deep learning based smartphone application for early detection of nasopharyngeal carcinoma using endoscopic images
- Tracking temporal progression of benign bone tumors through X-ray based detection and segmentation
- Clinical application research on the titanium metal metatarsal prosthesis designed through FEA and manufactured by 3D printing