sCIN: a contrastive learning framework for single-cell multi-omics data integration
Bioinformatics Multi-omics IntegrationPosted by mhb on 2025-11-12 09:42:49 |
Share: Facebook | Twitter | Whatsapp | Linkedin Visits: 6374
Introduction
Single-cell multi-omics technologies (like scRNA-seq and scATAC-seq) provide rich insights into cellular heterogeneity but integrating data across modalities is challenging because of their different feature spaces and distributions. Existing methods often fail to capture nonlinear relationships or remove batch effects effectively. To address this, the authors introduce sCIN (single-cell Contrastive INtegration) — a neural network framework using contrastive learning to align multiple omics datasets into a shared latent space while preserving biological meaning.
Methods
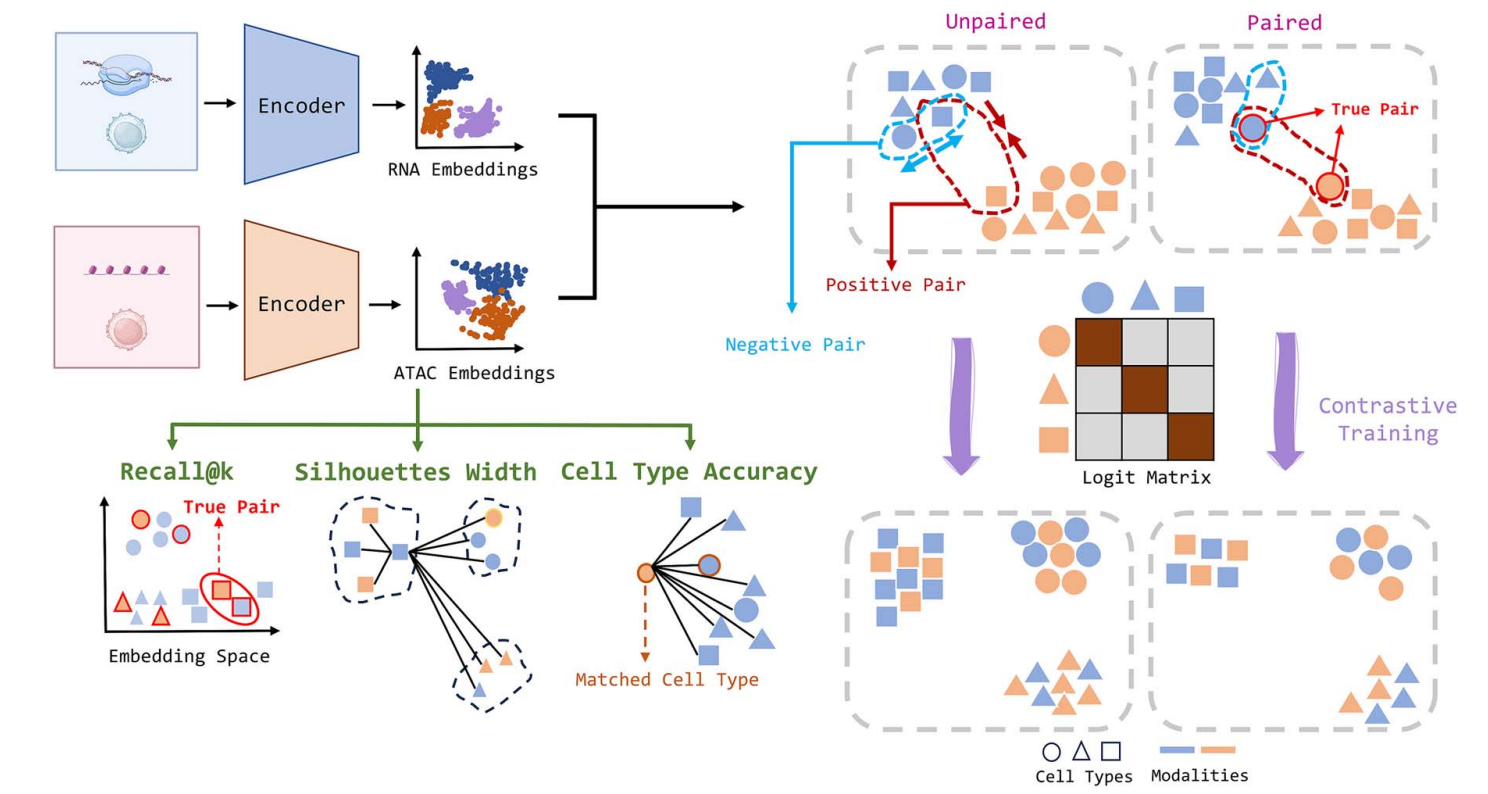
sCIN employs two modality-specific neural encoders that map data from different omics (e.g., RNA, ATAC, or ADT) into a shared low-dimensional space. It applies a contrastive loss to bring embeddings of similar cells (positive pairs) closer and push apart dissimilar ones (negative pairs).
-
Paired data: True cell pairs across modalities are positive.
-
Unpaired data: Cells with the same type across modalities are positive.
The framework was evaluated using multiple datasets (SHARE-seq, PBMC, CITE-seq, and Muto-2021) and compared against state-of-the-art methods like scGLUE, scBridge, sciCAN, MOFA+, Con-AAE, and Harmony using metrics such as Recall@k, ASW, cell type accuracy, and median rank.
Results
Across both paired and unpaired datasets, sCIN achieved:
-
The highest Recall@k and lowest median rank, indicating superior integration of modalities.
-
Better clustering (ASW > 0.6) and cell-type accuracy (~0.7–0.85), outperforming existing models.
-
In unpaired settings, sCIN still effectively aligned modalities even with small overlap between datasets, maintaining high biological consistency.
Conclusion
sCIN is a robust and flexible framework for integrating multi-omics single-cell data. It surpasses other methods in maintaining biological relevance while eliminating technology-driven variation. Its design avoids overfitting by using all matched pairs without hard mining.
However, it relies on accurate cell-type labels and currently supports only two modalities; future work aims to extend it to label-free and multi-modality integration.
Search
Categories
Recent News
- Clinical application research on the titanium metal metatarsal prosthesis designed through FEA and manufactured by 3D printing
- Manually weighted taxonomy classifiers improve species-specific rumen microbiome analysis compared to unweighted or average weighted taxonomy classifiers
- A deep learning based smartphone application for early detection of nasopharyngeal carcinoma using endoscopic images
- sCIN: a contrastive learning framework for single-cell multi-omics data integration
- CustOmics: A versatile deep-learning based strategy for multi-omics integration
- Tracking temporal progression of benign bone tumors through X-ray based detection and segmentation
- mCNN-GenEfflux: enhanced predicting Efflux protein and their super families by using generative proteins combined with multiple windows convolution neural networks
- Evaluation of normalized T1 signal intensity obtained using an automated segmentation model in lower leg MRI as a potential imaging biomarker in Charcot– Marie–Tooth disease type 1A
Popular Paper
- sCIN: a contrastive learning framework for single-cell multi-omics data integration
- CustOmics: A versatile deep-learning based strategy for multi-omics integration
- A deep learning based smartphone application for early detection of nasopharyngeal carcinoma using endoscopic images
- Tracking temporal progression of benign bone tumors through X-ray based detection and segmentation
- Clinical application research on the titanium metal metatarsal prosthesis designed through FEA and manufactured by 3D printing